
Every cell in your body contains strands of DNA wrapped into tightly bound structures called chromosomes. The DNA in your cells contains genetic information, encoded in the form a linear sequence of nucleotide bases. These sequences of bases contain the instructions for the construction of proteins. Molecules of RNA extract the information from DNA and, along with ribosomes, use that information to construct the proteins specified by the genetic code. This process is called translation.
In the context of genetics, a stop codon is a nucleotide triplet in messenger RNA that signifies the termination of protein translation. Essentially, a stop codon is a specific cluster of nucleotides that tells protein construction mechanisms to stop chaining amino acids into a polypeptide chain. Stop codons are the bits of information that tell the body “Hey! This protein is finished!” Stop codons work by initiating the release of release factors, proteins that disassociate the ribosomal subunits and free the polypeptide chain.
In the human genetic code, there have been identified 3 stop codons, each represented as a triplet of nucleotide bases.
In RNA:
- UAG (“amber”)
- UAA (“opal”)
- UGA (“ochre”)
In DNA:
- TAG
- TAA
- TGA
The three stop codons appear differently in DNA and RNA because RNA contains the U base in place of the T base in DNA.
Stop codons are paired with “start codons” that tell the cellular machinery the beginning of a DNA sequence that specifies a specific protein. Without start or stop codons, the mechanisms that read DNA and RNA would not know where to start and when to finish constructing proteins. The most common start codon is the nucleotide triplet AUG (ATG in DNA). Almost every eukaryotic organism uses the triplet AUG as a start codon.
What Are Codons?
How are proteins made in cells? Proteins are made by reading genes which are encoded in the form of sequences of nucleotide bases in DNA. The process of going from DNA to protein is called gene expression. In the first step of gene expression, a specific gene is “rewritten” in the form of mRNA. This process is called transcription. After transcription, the mRNA is “decoded” to build the proteins. This is called translation.
During translation information encoded in mRNA is read to build proteins. Specifically, the genetic code in mRNA specifies a specific polypeptide chain consisting of a determinate sequence of amino acids. The exact order of amino acids is specified by the order of the nucleotide bases in mRNA. The instructions in mRNA take the form of triplets of RNA bases (A, C, U, G) called codons. Each specific codon specifies a particular amino acid. For example, the codon GAG specifies glutamate and the codon ACG specifies threonine. So, if the mRNA contains a GAG codon then an ACG codon, the body will create a linear chain beginning with glutamate and then threonine. The exact order of amino acids determines the shape and function of the protein. All in all, there are 61 distinct codons in the human genome that encode for 20 amino acids.

A table specifying the codons and the respective amino acids they signify. Credit: WikiCommons CC0 1.0
There is also a special codon called a start codon that signifies the beginning of the polypeptide chain. By far the most common start codon found in eukaryotes is codon AUG. AUG is also used to specify methionine.
There are three other codons that do not specify amino acids. These are called stop codons and signify when a protein is complete. The most common stop codons are UAA, UGA, and UAG, though a handful of different stop codons are used in some organisms.
To understand the important role stop codons play, consider what would happen if there weren’t any stop codons. Imagine a hypothetical gene containing the codons CCU, GCU, and CAU. These three codons specify proline, alanine, and histidine, respectively. Reading through these codons, the cell would attach one proline to one alanine to one histidine. Without a stop codon, the body would continue to read the mRNA and keep attaching amino acids. The proteins would then be nonfunctional because they are too large. Stop codons tell cellular machinery that a particular protein is done being constructed.
How, Exactly, Do Codons Work?
So far, we have learned that codons are particular triplets of nucleotides that specify sequences of amino acids. The code in mRNA is read and a linear chain of amino acids is constructed according to the order of codons in the mRNA. Stop codons are special codons that tell the body to stop protein translation. How, exactly, do codons specify amino acids though? That is, what are the molecular mechanisms by which codons perform their function?
The answer: tRNA. tRNA (transfer RNA) is a special kind of RNA molecule that serves as the bridge between the codons in mRNA and the amino acids they specify. On one end of tRNA is a specific anticodon; that is, a specific triplet of nucleotides that are the complementary pairs of the bases found in mRNA, and on the other end is the specified amino acid. tRNA binds to mRNA via codon-anticodon bonds, bringing along the specified amino acid.
For instance, say an mRNA strand has the codon UCA (serine). The appropriate tRNA contains the anticodon AGU and the amino acid serine. The tRNA anticodon will bind to the mRNA codon and bring along with it the serine molecule. There are several different types of tRNA each that read one or a few codons. The carboxyl group of the amino acid is then joined to the amine group of another amino acid via a hydrolysis reaction.
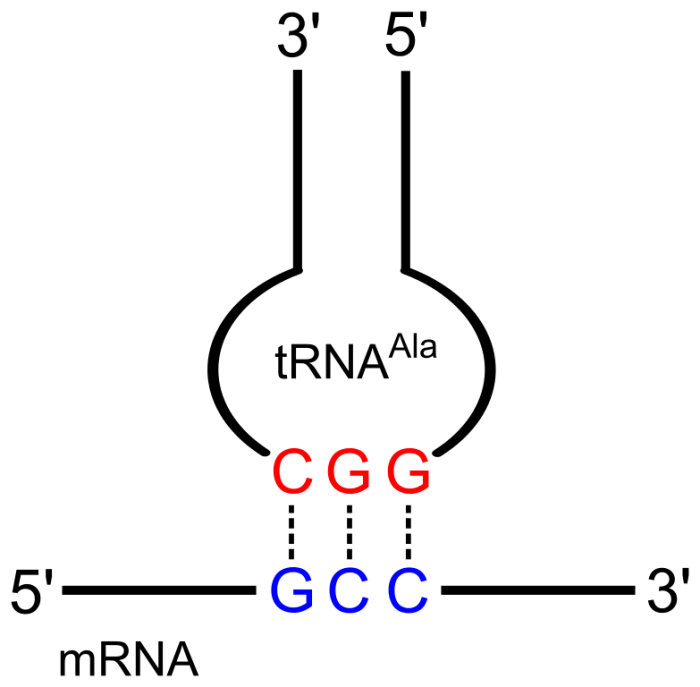
A codon (blue) anticodon (red) pair. Credit: Yikrazuul via WikiCommons CC BY-SA 3.0
This is not the full story though. mRNA and tRNA by themselves cannot build proteins. Ribosomes are organelles where the polypeptide chain is actually constructed. Ribosomes are made out of special kind of RNA called ribosomal RNA (rRNA) and several distinct proteins. Ribosomes consist of two parts: a large unit and a small unit that enclose mRNA, kind of like the two pieces of bread on a sandwich.
Ribosomes contain small “slots” that help tRNA find their matching codons. There are three places for tRNA to bind to ribosomes, called the A, P, and E slots. Ribosomes are important because they organize mRNA and tRNA to make sure proteins are constructed correctly. Aside from serving as the physical scaffolding for protein construction, ribosomes also have enzymes that catalyze the hydrolysis reaction that joins amino acids into a polypeptide chain.
Stop Codons
What about stop codons though? As stated previously, stop codons do not encode for amino acids, so how do those work? Instead of coding for tRNA and amino acids, stop codons are recognized by proteins called release factors. Release factors cause the ribosomal subunits to dissociate, freeing the polypeptide chain. which can go on to post-translation processing and modification. Currently, the exact molecular mechanisms behind the recognition of stop codons by release factors are not well understood. There are no known tRNA subunits that have stop codon anticodons, so it does not seem as if termination is directly caused by other tRNA molecules. There is some evidence that ribosomal RNA may play some role in recognizing stop codons in mRNA but so far there is no conclusive evidence.
Sometimes stop codons can pop up as the result of a random mutation. So-called “nonsense mutations” can introduce premature stop codons into the genome, which will cause early termination of translation. This is partially the reason why the DNA of so many organisms has evolved to have several redundant bits of code, so that if one fails others can take up the mantle.
Alternatively, sometimes DNA can mutate to get rid of stop codons. In these cases translational mechanisms do not get the signal to stop making proteins, so they will translate regions that are meant to be left untranslated, or are part of a separate gene sequence. Most proteins formed from a gene with “non-stop” mutations are nonfunctional because they are extremely long.
Additionally, “hidden stops” are when a non-stop codon is read as a stop codon because the RNA reading mechanisms become shifted one place to the right or left. For example, say an mRNA strand has the sequence GUCUUAGAC. Normally, the codons would be read in triplets, like GUC, UUA, and GAC. introducing a frameshift +1 to the right cuts off the leftmost base and the RNA would be read as UCU, UAG, GAC. Moving the frame to the right. The result is that the non-stop codon gets read as a stop codon. This can happen in normal segments of mRNA too, leading the body to create nonsense proteins that do not serve any function.
As a summary, codons are sequences of nucleotide bases in mRNA that signify specific amino acids. Stop codons are a special class of codons that do not code for amino acids, but instead signal to translation mechanisms to stop making proteins. Stop codons tell the body where one gene ends and when to stop chaining amino acids into polypeptide chains.








