An explosion of research in recent years has brought to light the multitude of ways in which the gut microbiota influences animal health and development. The gut microbiota is incredibly diverse, comprising hundreds of species, and varies both across and within individuals over time. While scientists can identify who is present in the microbiota, how and/or why particular microbes are present is largely unknown. In particular, it is not clear how microbes adapt to their hosts, nor is it known what microbial traits are important for host colonization.
Evolutionary adaptation is a process by which organisms become better suited to a particular environment or condition. In short, individuals in populations naturally vary in their traits, and these traits can sometimes be advantageous — those individuals are more likely to survive and reproduce. With every generation, these lineages expand, increasing the overall success of the evolving population. Due to the generally large population sizes and short generation times of bacteria, adaptation can occur very quickly. So quickly, in fact, that we can use evolution as a tool in the laboratory to observe adaptation in real time and identify which bacterial traits are advantageous under particular conditions.
Animal models allow us to study and manipulate host-microbe systems that we couldn’t otherwise control in natural settings. The zebrafish is a great example of such a model. These animals can be raised in the absence of microbes, allowing the researcher to expose the animals to only particular microbial symbionts of interest, and track them throughout the colonization process. Moreover, the zebrafish model system captures important ecological features of natural host-microbe systems. For example, the aquatic environment of the fish serves as a medium through which microbes can move from fish to fish. In addition, because fish produce many progeny, we can easily rear them in large groups, simulating natural populations of animals.
We experimentally evolved a bacterial strain (a common microbe from fish known as Aeromonas) to optimize its ability to colonize the gut of the larval fish. To do this, we used “serial passage,” a technique that involves inoculating groups of fish with microbes derived from previous groups. Only Aeromonas populations present in the intestines of the fish were used to inoculate new groups of zebrafish. After each inoculation, the microbes increased in number and occasionally mutants arose that were better at colonizing the larval fish. These mutants were then passed on to the next generation of fish, thereby enriching for natural variants that are good at colonizing the gut (Figure 1).
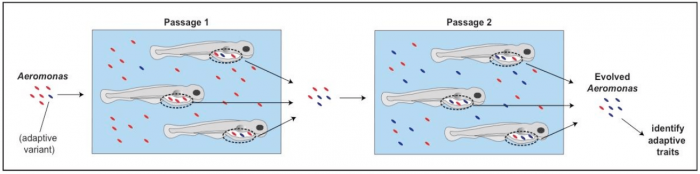
Figure 1. Schematic of experimental evolution of Aeromonas to the zebrafish gut. Aeromonas was serially passaged in larval zebrafish, each time selecting only gut-associated Aeromonas populations to inoculate the next group of fish. Here, only two passages are depicted, but in our actual experiments, there were 22 passages. The blue Aeromonas represent a natural variant that has a trait that is advantageous for gut colonization. With each passage, it expands within the evolving population. The goal of the project was to characterize the adaptive traits of the evolved isolates. Note: zebrafish and bacteria are not to scale. Image courtesy Cathy Robinson.
Next, we asked HOW these variants were better at colonizing the gut. Generally, it’s assumed that microbes adapt to animal guts by acquiring traits that impact their growth or persistence specifically within the gut. Surprisingly, we found that the first variants of Aeromonas enriched in our experiment did not have gut-specific adaptations, but instead acquired the ability to immigrate into the fish from the aquatic environment more quickly than the ancestral strain. This provides an advantage both for initial colonization and also throughout colonization as space becomes available. Because they are able to get to the fish gut faster and begin growing, they become a larger proportion of the final gut population. It was only later in the experiment that we observed variants with gut-specific adaptations. In essence, we found that Aeromonas first evolved to more efficiently get into the host, and secondarily adapted to the conditions within the host.
This work provides insight into the adaptive strategies of animal gut microbes. It also contributes to a growing body of evidence supporting the role of transmission in shaping host-associated microbial communities. Finally, it demonstrates that host-microbe systems should be studied with a keen consideration of the broader ecological context inherent in these complex ecosystems. Better understanding what controls the membership of gut microbiota will one day help us develop approaches to treat or prevent microbiota-associated diseases.
These findings are described in the article entitled Experimental bacterial adaptation to the zebrafish gut reveals a primary role for immigration, recently published in the journal PLOS Biology.









