
Primary immunodeficiencies (PID) are a heterogeneous group of disorders that disturb the host’s immunity, creating susceptibility to infections. PIDs are genetically diverse, with mutations in many different genes capable of causing immunodeficiency. The clinical symptoms of PIDs include, but are not limited to, susceptibility to infections, inflammation, and autoimmunity, although each gene mutated, and indeed each individual mutation, can lead to different manifestations.
Central to understanding PIDs is to understand which immune cell type is rendered defective by the mutation the patient carries. The type of infections the patient develops is often a key indicator of the underlying immunodeficiency; for example, pulmonary infections and bacterial septicemia are associated with B cell defect, whereas fungal susceptibility is associated with defects in certain types of T cells. Candidate pathways can be investigated using genetics and immune screening, and successful identification of the underlying causes allows a treatment program to be tailored to the patient.
Roifman syndrome is a rare autosomal recessive PID. It was originally believed to be an X-linked syndrome since the first six patients to be described were male. It was recently discovered, however, that the disease is not X-linked and is caused by mutations in the small nuclear RNA noncoding gene RNU4ATAC. To date, only ten cases from seven families have been reported with Roifman syndrome, although unreporting is expected, with most clinicians unaware of the disease. Roifman patients present with growth retardation, cognitive delay, retinal dystrophy, and spondyloepyphyseal dysplasia. Roifman patients are also immunodeficient, only presenting with recurrent respiratory infections, otitides, and a diminished vaccine response. In a recent study, we sought to understand the nature of the immunodeficiency suffered by Roifman patients, to allow better treatment of this devastating syndrome.
In a study led by Prof Adrian Liston from the VIB in Belgium, Roifman syndrome patients were extensively studied at the immunological and molecular level. By investigating the immune system of the Roifman patients, the authors found that patients exhibited a major problem in the white blood cell type known as B cells. Dr. Josselyn Garcia-Perez likens the defect to the B cells being trapped in childhood. “B cells are born in the bone marrow and migrate into the blood. These ‘transitional’ B cells, fresh out of the bone-marrow are like children and still need to mature before they can do the full workload of adult B cells. In Roifman patients, their B cells get stuck at this transitional stage, and the patients never get a normal set of B cells.” In the absence of mature B cells, the patients are unable to make antibodies in response to infections, leading to prolonged infections and frequent reinfection.
The authors further investigated this defect in B cells at the molecular level. The patients bore mutations in the gene RNU4ATAC, which is an essential part of the RNA splicing complex. RNA is often described as the messenger that links the instructions encoded in DNA to the protein building blocks that make up the cell. RNA is generated by copying small lengths of stable, and largely inert, DNA into the dynamic and unstable RNA molecule. This RNA is then able to leave the nucleus of the cell and travel to molecular factories called ribosomes that read the instructions encoded in RNA to make proteins.
The process of copying instructions from DNA to RNA is complicated by every set of instructions in the DNA being divided into multiple different sites. Professor Liston describes it as, “between each page of the instruction manual is many pages of gibberish. The RNA message needs to have this gibberish cut out in order for the instructions to come together into a single message.”
This process is known as splicing and involves piecing together the key code of the gene (or “exons”) into a single long strip of sequence. There are two sets of machinery that can splice RNA in the cell – the major splicing complex and the minor splicing complex. The major splicing complex is so essential to the cell that defects in this process are lethal at the single cell stage and never observed in patients. The minor splicing complex is less important, as only a handful of genes use it to piece together their instructions. The RNU4ATAC gene that is mutated in Roifman patients is part of this minor splicing complex, with the symptoms of the disease being driven by errors introduced in the genes that need it.
By purifying the transitional B cells that were defective in Roifman patients, the authors were able to use a next-generation sequence to find out which genes were incorrectly edited due to the mutation in the minor splicing complex. Molecular analysis identified the gene MAPK1 as incorrectly spliced in Roifman patients. Dr. Garcia-Perez explains, “Essentially, Roifman patient B cells were leaving in one page of nonsense in the RNA instructions to make the protein MAPK1, creating a functional problem in this essential gene.”
The problem in MAPK1 resulted in transitional B cells that were unable to receive signals from the growth factor BAFF. Without these growth signals, the transitional B cells were not able to mature into the fully functional B cells needed to make antibodies. The authors hope that these findings will result in improved treatment for Roifman patients. Prof Liston says, “The nature of this problem suggests that the immune issues in Roifman patients could be solved by bone-marrow transplantation. Also, now that we know the problem the B cells have in hearing the receiving BAFF signal, future studies could test whether giving extra BAFF to the patients rescues the B cell defect.”
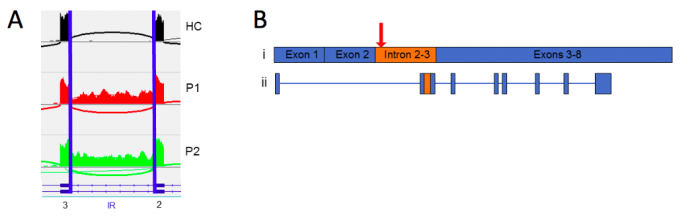
Figure 1. Splicing defect in MAPK1. (A) Sashimi plot showing MAPK1 splicing in healthy controls (HC) and Roifman patients (P1 and P2). The “IR” region is edited out successfully in the healthy control but is often left in Roifman patients. (B) The splicing error in MAPK1 results in an extra sequence (orange) being left inside the correct (blue) RNA sequence. This defective RNA molecule fails to produce the protein MAPK1. Image published with permission from Elsevier from https://doi.org/10.1016/j.jaci.2017.11.061.
These findings are described in the article entitled Abnormal differentiation of B cells and megakaryocytes in patients with Roifman syndrome, recently published in the Journal of Allergy and Clinical Immunology. This work was conducted by Jessica Heremans, Josselyn E. Garcia-Perez, Ernest Turro, Susan M. Schlenner, Ingele Casteels, Roxanne Collin, Francis de Zegher, Daniel Greene, Stephanie Humblet-Baron, Sylvie Lesage, Patrick Matthys, Christopher J. Penkett, Karen Put, Kathleen Stirrups, on behalf of the National Institute for Health Research BioResource, Chantal Thys, Chris Van Geet, Erika Van Nieuwenhove, Carine Wouters, Isabelle Meyts, Kathleen Freson, and Adrian Liston.









