
In order to survive in an environment populated by hostile viruses ready to parasite them, mammalian cells have evolved a sophisticated variety of constitutively expressed receptors to detect extra- or intracellular pathogen-associated molecular signatures.
Upon activation by binding to their targets, such receptors trigger a specific intracellular signaling which leads to a potent production and secretion of antiviral molecules in the surrounding environment. Such secreted factors named “interferons” not only strengthen the antiviral defenses of nearby uninfected cells but also act as chemical messengers to activate and recruit more specialized components of the immune system1,2.
In addition to their sensing compartment, mammalian cells are also equipped with a heterogeneous arsenal of “restriction factors,” germ-line-encoded proteins that actively antagonize viral replication throughout a plethora of different mechanisms3. Unlike the immune response that needs its time in order to become operational, restriction factors are already present at the moment of the infection, thereby forming our very first line of defense. Their expression is very often upregulated by interferons and their presence in a specific cell type represent a severe obstacle for virus replication, forcing the latter ones to constantly evolve new ways to neutralize or evade them4,5.
The human PYHIN family consists of four members: γ- interferon-inducible protein 16 (IFI6), myeloid cell nuclear differentiation antigen (MNDA), interferon-inducible protein X (IFIX) and absent in melanoma 2 (AIM2). As represented in Figure 1, all the PYHIN proteins harbor a PYRIN domain (PYD), which mediates the interactions with other PYD-containing proteins, and one or more HIN domains, required for DNA binding6. The presence or absence of multiple nuclear localization signals (NLS) dictates the cellular compartmentalization of these proteins: in physiological conditions, IFI167, MNDA8,9 and IFIX10 exhibit a nuclear localization while AIM2 is located in the cytoplasm11,12.
The PYHIN proteins seem to be involved in regulating several cellular processes, including proliferation, differentiation, transcription, DNA damage, and apoptosis13. Given though their ability to bind exogenous DNA in a sequence-independent manner7,11,14–16, emerging evidence strongly supports the idea that the PYHIN proteins play a central role in activating the intrinsic and innate response upon DNA virus infection. What I am currently investigating during my Ph.D. is the mechanism through which these proteins can suppress HIV replication.
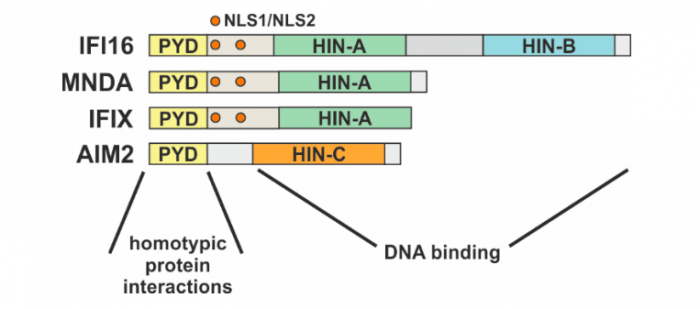
Figure 1: Schematic representation of the human PYHIN family, with highlighted the different functional regions. Credit: Matteo Bosso
HIV belongs to the family of Retroviridae due to the presence of two unique enzymes that are not found in any other living being: (i) a reverse transcriptase, catalyzing the conversion of the single-stranded viral RNA into a double-stranded DNA molecule and (ii) an integrase, responsible for the stable insertion of the viral DNA into our genome. Reverse transcription gives rise to several intermediate products (RNA:DNA hybrids, single stranded and double stranded DNA)17 and defects during this process could lead to an accumulation of viral DNA fragments susceptible for immune sensing18.
Jakobsen et al. first demonstrated that IFI16 binds to single-stranded DNA species derived from HIV in human macrophages19 and a follow-up study revealed that this protein cooperates with another DNA sensor (cGAS – cyclic dinucleotide GMP-AMP synthase) to induce the interferon response20. This synergistic behavior between IFI16 and cGAS seems not restricted exclusively to HIV, as it has been previously observed in other cell types21 as well as in the presence of other pathogens21,22. Interestingly, the IFI16-mediated sensing of viral DNA is also associated with the progressive loss of CD4+ T cells, a hallmark of AIDS development, as it promotes cell death upon abortive HIV infection23. In addition to this, our group identified IFI16 as a potential HIV restriction factor during a genome-wide screening and our previously published data suggest that its antiviral activity is independent of interferon activation24.
Amongst the PYHIN proteins, IFI16 is the one that received the highest attention, as it was the first DNA sensor to act within the nucleus25 — challenging the dogma that detection of viral nucleic acids strictly depends on the cellular compartment. One study identified a crucial mechanism through which cells carefully regulate the distribution of IFI1626 although it is not fully understood what regulates its translocation upon DNA sensing.
Furthermore, it is also not clear which stimuli dictates the activation of either IFI16 or AIM2 in the presence of cytosolic DNA. While several pathogenic bacteria were shown to trigger AIM2 response27, recent studies revealed that also many DNA viruses induce its activation28-30, suggesting a synergistic sensing between IFI16 and AIM2. Finally, also IFIX emerged as an exogenous-DNA sensor13 even though the precise mechanisms through which it stimulates the interferon response are still unknown.
Besides their role as immune sensors, there is increasing evidence supporting the idea that the nuclear PYHIN proteins can directly suppress viral replication. IFI16 was shown to antagonize a broad spectrum of DNA viruses ranging from Herpes31 to Papilloma32 although less information is available concerning IFIX33 and MNDA34. Even though the precise mechanisms of action are still a matter of discussion, the general hypothesis is that the PYHIN proteins shut down the expression of viral genes by reducing the recruitment of cellular complexes (like the RNA polymerase II). Given their role in controlling viral replication, are therefore not surprising the findings that some viruses have evolved peculiar ways to counteract or hijack these restriction factors.25,33,35-37
With the help of Dr. Dominik Hotter and Junior Professor Daniel Sauter, and the supervision of Professor Frank Kirchhoff, it would be our task to find out whether the human PYHIN proteins can exert the same antiviral function against HIV and if — and how — the virus found a way to circumvent their presence in its target host cells.
This article is based on findings conducted by Matteo Bosso from the Institute of Molecular Virology, Ulm, Germany.
References:
- Komatsu, T., Nagata, K. & Wodrich, H. The role of nuclear antiviral factors against invading DNA viruses: The immediate fate of incoming viral genomes. Viruses (2016). doi:10.3390/v8100290
- Chan, Y. K. & Gack, M. U. Viral evasion of intracellular DNA and RNA sensing. Nat. Rev. Microbiol. (2016). doi:10.1038/nrmicro.2016.45
- Bieniasz, P. D. Intrinsic immunity : a front-line defense against viral attack. 5, 1109–1115 (2004).
- Harris, R. S., Hultquist, J. F. & Evans, D. T. The restriction factors of human immunodeficiency virus. Journal of Biological Chemistry (2012). doi:10.1074/jbc.R112.416925
- Sauter, D. & Kirchhoff, F. Multilayered and versatile inhibition of cellular antiviral factors by HIV and SIV accessory proteins. Cytokine Growth Factor Rev. (2018). doi:10.1016/j.cytogfr.2018.02.005
- Connolly, D. J. & Bowie, A. G. The emerging role of human PYHIN proteins in innate immunity: Implications for health and disease. Biochemical Pharmacology (2014). doi:10.1016/j.bcp.2014.08.031
- Unterholzner, L. et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. (2010). doi:10.1038/ni.1932
- Goldberger, A., Brewer, G. & Hnilicat, L. S. Nonhistone Protein Antigen Profiles of Five Leukemic Cell Lines Reflect the Extent of Myeloid Differentiation. Blood 63, 701–710 (1984).
- Goldberger, A., Hnilicatll, L. S., Caseyli, S. B. & Briggssli, R. C. Properties of a Nuclear Protein Marker of Human Myeloid Cell Differentiation*. J. Biol. Chem. 261, 4726–4731 (1986).
- Ding, Y. et al. Antitumor activity of IFIX, a novel interferon-inducible HIN-200 gene, in breast cancer. Oncogene 23, 4556–4566 (2004).
- Hornung, V. et al. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature (2009). doi:10.1038/nature07725
- Khare, S. et al. The PYRIN domain-only protein POP3 inhibits AIM2-like receptor inflammasomes and regulates responses to DNA virus infections. 15, 343–353 (2014).
- Diner, B. A. et al. The functional interactome of PYHIN immune regulators reveals IFIX is a sensor of viral DNA. Mol. Syst. Biol. 11, 787 (2015).
- Fernandes-Alnemri, T., Yu, J., Datta, P., Wu, J. & Alnemri, E. S. AIM2 activates the inflammasome and cell death in response to cytoplasmic DNA. Nature 458, 509–513 (2009).
- Bürckstümmer, T. et al. An orthogonal proteomic-genomic screen identifies AIM2 as a cytoplasmic DNA sensor for the inflammasome. 10, (2009).
- Roberts, T. L. et al. HIN-200 Proteins Regulate Caspase Activation in Response to Foreign Cytoplasmic DNA. Science (80-. ). 115, 1057–1061 (2009).
- Abbink, T. E. M. & Berkhout, B. HIV-1 reverse transcription initiation : A potential target for novel antivirals ? Virus Res. 134, 4–18 (2008).
- Doitsh, G. et al. Abortive HIV Infection Mediates CD4 T-Cell Depletion and Inflammation in Human Lymphoid Tissue. Cell 143, 789–801 (2011).
- Jakobsen, M. R. et al. IFI16 senses DNA forms of the lentiviral replication cycle and controls HIV-1 replication. Proc. Natl. Acad. Sci. U. S. A. 110, E4571-80 (2013).
- Jønsson, K. L. et al. IFI16 is required for DNA sensing in human macrophages by promoting production and function of cGAMP. Nat. Commun. (2017). doi:10.1038/ncomms14391
- Almine, J. F. et al. IFI16 and cGAS cooperate in the activation of STING during DNA sensing in human keratinocytes. Nat. Commun. (2017). doi:10.1038/ncomms14392
- Hansen, K. et al. Listeria monocytogenes induces IFN b expression through an IFI16- , cGAS- and STING-dependent pathway. EMBO J. 33, 1654–1666 (2014).
- Monroe, K. M. et al. IFI16 DNA Sensor Is Required for Death of Lymphoid CD4 T Cells Abortively Infected with HIV. Science (80-. ). (2014). doi:10.1126/science.1243640
- McLaren, P. J. et al. Identification of potential HIV restriction factors by combining evolutionary genomic signatures with functional analyses. Retrovirology (2015). doi:10.1186/s12977-015-0165-5
- Orzalli, M. H., DeLuca, N. A. & Knipe, D. M. Nuclear IFI16 induction of IRF-3 signaling during herpesviral infection and degradation of IFI16 by the viral ICP0 protein. Proc. Natl. Acad. Sci. (2012). doi:10.1073/pnas.1211302109
- Li, T., Diner, B. A., Chen, J. & Cristea, I. M. Acetylation modulates cellular distribution and DNA sensing ability of interferon-inducible protein IFI16. Proc. Natl. Acad. Sci. (2012). doi:10.1073/pnas.1203447109
- Man, S. M., Karki, R. & Kanneganti, T. D. AIM2 inflammasome in infection, cancer, and autoimmunity: Role in DNA sensing, inflammation, and innate immunity. European Journal of Immunology (2016). doi:10.1002/eji.201545839
- Huang, Y. et al. Interaction between HCMV pUL83 and human AIM2 disrupts the activation of the AIM2 inflammasome. doi:10.1186/s12985-016-0673-5
- Lugrin, J. & Martinon, F. The AIM2 inflammasome : Sensor of pathogens and cellular perturbations. 99–114 (2018). doi:10.1111/imr.12618
- Maruzuru, Y. et al. Herpes Simplex Virus 1 VP22 Inhibits AIM2- Dependent Inflammasome Activation to Enable Article Herpes Simplex Virus 1 VP22 Inhibits AIM2-Dependent Inflammasome Activation to Enable Efficient Viral Replication. Cell Host Microbe 23, 254–265 (2018).
- Dell’Oste, V. et al. The interferon-inducible DNA-sensor protein IFI16: A key player in the antiviral response. New Microbiologica (2015).
- Lo Cigno, I. et al. The Nuclear DNA Sensor IFI16 Acts as a Restriction Factor for Human Papillomavirus Replication through Epigenetic Modifications of the Viral Promoters. J. Virol. 89, 7506–20 (2015).
- Crow, M. S. & Cristea, I. M. Human antiviral protein IFIX suppresses viral gene expression during HSV-1 infection and is counteracted by virus-induced proteasomal degradation. 1–47 (2017).
- Liu, S.-Y., Sanchez, D. J., Aliyari, R., Lu, S. & Cheng, G. Systematic identification of type I and type II interferon-induced antiviral factors. Proc. Natl. Acad. Sci. 109, 4239–4244 (2012).
- Lanfranca, M. P., Mostafa, H. H. & Davido, D. J. HSV-1 ICP0: An E3 Ubiquitin Ligase That Counteracts Host Intrinsic and Innate Immunity. Cells 3, 438–454 (2014).
- Li, T., Chen, J. & Cristea, I. M. Human cytomegalovirus tegument protein pUL83 inhibits IFI16-mediated DNA sensing for immune evasion. Cell Host Microbe (2013). doi:10.1016/j.chom.2013.10.007
- Biolatti, M. et al. Regulatory Interaction between the Cellular Restriction Factor IFI16 and Viral pp65 (pUL83) Modulates Viral Gene Expression and IFI16 Protein Stability. J. Virol. 90, JVI.00923-16 (2016).









