
Cell shape (morphology) and its temporal variation (morphodynamics) are the phenotypic outcomes of many cellular processes, including migration, tissue organization, proliferation, differentiation, and death. Thus, they often serve as indicators of physiological and pathological states of cells.
The best examples, perhaps, are cytology and histology, where – despite all the advances in genomic and molecular profiling – pathologists still assess disease progression mostly based on cell and tissue morphological features. Moreover, innumerable molecular signaling pathways converge onto the regulation of the cytoskeleton to direct and determine morphological changes of cells. Therefore, quantification on cell morphology can also provide incisive information on signaling states of cells.
While many assays rely on global cell shape descriptors, such as cell area change and degree of polarization, the dynamics of shape changes is much less explored. In our recent study “Profiling cellular morphodynamics by spatiotemporal spectrum decomposition,” published in the journal PLOS Computational Biology, we propose a framework to profile cellular morphodynamics with high spatiotemporal resolution for probing signaling states of cells. The study relies on previous work by our lab to accurately track and quantify the movement of cells filmed by high-resolution microscopy.
Frequency consistency upon spontaneous protruding cells uncovered by Hilbert-Huang Transform
A population of spontaneously protruding fibroblast-like cells, even though genomically homogeneous, present vastly heterogeneous appearance and motility (See Fig. 1(a) and (b)). Some cells are active, while others are quiescent. In our study, we raised the first question whether there is an indicator or feature extracted from the cell edge motion that would reflect the homogeneity in genetic background, or at the molecular level, the consistency among cells in their wiring of regulatory circuitry.
By utilizing the Hilbert-Huang Transform (HHT), we can decompose the edge velocity time series at each a specific cell edge location into finite component signals, referred to as intrinsic mode functions (IMFs), and compute their instantaneous frequency and amplitude at each time point. Quite remarkably, we found the instantaneous frequency spectra among ~50 Cos7 fibroblast-like cells nearly invariant. Their marked differences in motility were fully captured by only the instantaneous magnitude spectra (See Fig. 1(c) and (d)).
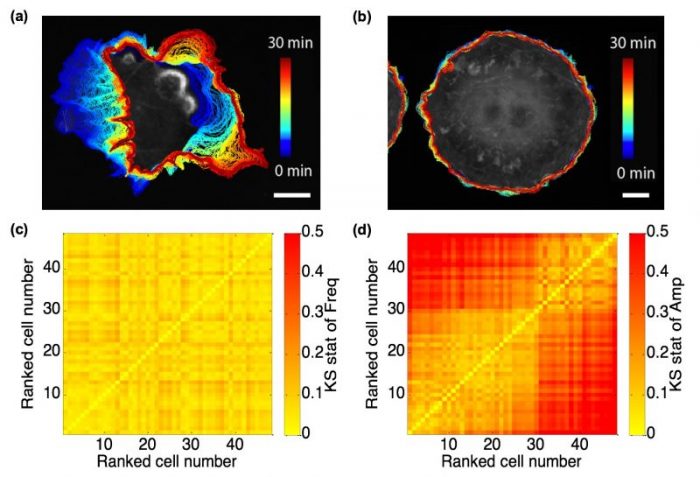
Fig. 1 Fibroblast-like cells with different motility, and comparison between frequency and amplitude spectra. (a) and (b) Cos7 cells representing active (a) and quiescent (b) motility with overlaid cell boundaries migrating from early (blue, 0 min) to late (red, 30 min) time points; scale bar = 20 μm. (c) and (d) Heatmaps of Kolmogorov-Smirnov (KS) statistics for instantaneous frequency (c) and amplitude (d) distributions among 48 spontaneously protruding Cos7 cells. The sequence of cells in the heatmap is defined in ascending order of the mean velocity over all cell edge locations and time points. Republished from the PLOS article: https://doi.org/10.1371/journal.pcbi.1006321
Acutely perturbed Vav2 signaling leads to shift in the instantaneous frequency spectra
The conservation of instantaneous frequency spectra in spontaneously migrating Cos7 cells inspired us to ask whether induced changes in the wiring of morphogenetic signaling pathways would affect the frequency distribution or whether such changes would also be reflected by mere shifts in the instantaneous frequency spectra.
To test this we employed an optogenetic construct developed in Klaus Hahn’s lab at University of North Carolina at Chapel Hill, to acutely photo-inhibit the guanosine exchange factor (GEF) Vav2, which is a known signaling component in cell morphogenic pathways. Indeed, with gradually stronger inhibition of the factor, we did observe a shift of the frequency spectra. This indicated to us that the frequency spectra encode the signaling topology whereas the amplitude spectra encode the activation level of morphogenetic signals.
Spectral features define motion regimens along cell periphery
After analyzing and comparing the overall morphogenetic signaling states, we then wondered whether instantaneous frequency and amplitude spectra could also be employed to distinguish transient signaling states in localized subcellular regions.
To address this, we first manually selected subcellular regions of Cos7 cells that represent potent versus weak migrating polarity. In highly polar cells, front and back clearly showed significant differences in the frequency spectra, confirming that cell polarization implicates the stabilization of distinct signaling topologies over short distances. Based on this finding, we employed an unbiased clustering method – statistical region merging (SRM) – to identify transient spatiotemporal zones with distinct motion behavior.
Application of morphodynamic profiling to define distinct signaling zones
Finally, we show directly that the morphodynamically defined spatiotemporal zones correspond to different signaling regimes. To this end, we adopted a Förster resonance energy transfer (FRET) biosensor to probe with subcellular resolution the activation of the GTPase Rac1, which is the downstream effector of Vav2. We applied SRM to identify several edge motion zones and showed that distinct instantaneous frequency spectra correspond to distinct levels of Rac1 activation, as well as distinct levels of coordination between motion and signaling cycles.
Concluding remarks and future research
We propose a framework in this study to profile cellular morphodynamics using HHT (spectral decomposition + instantaneous frequency/amplitude computation) and SRM (unsupervised clustering). Apart from global spectral decomposition methods such as Fourier Transform, the HHT based decomposition captures oscillatory variations of cell edge motion between cell locations and over time. Our experiments then show that these motion regimens are directly linked to the local organization of signaling regimes. Moving forward, we intend to detect in a rigorous manner transient “signaling microdomains” that entail consistent regulatory signaling activities in subcellular regions. Moreover, the spectral decomposition and clustering may also be employed in the analysis of biosensor fluctuation signals, which may enable the complete mapping of subcellular regimes with a distinct regulation of signaling.
These findings are described in the article entitled Profiling cellular morphodynamics by spatiotemporal spectrum decomposition, recently published in the journal PLOS Computational Biology. This work was conducted by a team including Xiao Ma and Gaudenz Danuser from the University of Texas Southwestern Medical Center, and Onur Dagliyan and Klaus M. Hahn from the University of North Carolina at Chapel Hill.








