
The most beautiful, complicated and mystifying events in nature exist on a scale that we cannot directly witness. This is evident in the biological sciences, where living organisms can be only microns in size, and where standard microscopes are not sufficient to observe the details present at this scale. Harmonious to the need for advanced imaging, microscope technology has progressed immensely in the past few decades and has provided fuel for curiosity by allowing scientists to observe the microscopic world in more detail.
Techniques like electron microscopy allow the magnification of samples several thousand times, providing accurate visual accounts that are typically acceptable as supporting evidence of the items/processes under observation. It would be correct to assume that the impressive equipment associated with these technologies are usually only sustainable at high-level institutions that are capable of financing the maintenance and expertise required to accommodate these technologies.
An added downside to electron microscopy and similar imaging technologies are the costs and procedures involved with preparing samples to be viewed with these microscopes. They are characteristically tedious, time-consuming, and involve reagents and secondary equipment that are expensive and often dangerous.
To reduce the complexity and expense associated with conventional biological sample preparation for electron microscopy, we developed a protocol that eliminates the fixative Osmium tetraoxide (OsO4) and critical point drying, both expensive components of conventional sample preparation procedures.
To show that omitting these aspects of sample preparation is feasible, we visualized the interactions of an E. coli strain engineered to delivery DNA to mammals, with human cancer cells. Our results showed that basic morphology and interactions of the bacteria and human cells were preserved enough to produce satisfactory images even when using attenuated sample preparation procedures (Figure 1). Utilizing this affordable approach, we captured changes to the patterns of invasive bacteria-human cell adherence induced using Lipofectamine and PULSin, commercial cationic lipids that are routinely used for DNA and protein transfection respectively.
The behavior of this E. coli vector when interacting with human cells, especially its efficacy of attachment to the cell membrane translates directly to the success of eventual gene transfer from the vector to the host. We were able to manipulate these interactions using the aforementioned cationic lipids and observe significant changes in how the vector binds to the cell membrane of the host cells (Figure 2).
Omitting sample fixation with OsO4 and critical point /freeze-drying during preparation for electron microscopy reduced the preparation time and experimental cost of the procedure several-fold, and nonetheless yielded accurate representations of the biological interactions in question, with decent quality. We found mainly that thorough sample fixation using paraformaldehyde or glutaraldehyde, and gentle but precise dehydration of samples in ethanol gradients were the critical steps to produce samples of comparable integrity to those prepared using conventional high-end reagents and equipment.
The protocol we have highlighted represents a quick and affordable way to prepare biological samples for electron microscopy of basic structures in biological specimen without sacrificing pertinent morphological details. While this procedure may not be universally applicable to all biological specimen, it may be applied to most cultured mammalian and bacterial cells and may be useful routinely where advanced microscopic analysis of specimen is otherwise abandoned due to the intricacy and expense traditional of sample preparation.

Figure 1: Electron microscope micrograph obtained using simplified sample preparation, showing an HT1080 fibroblast (brown) being invaded by an engineered E. coli vector (Blue). Credit: Kumaran Narayanan
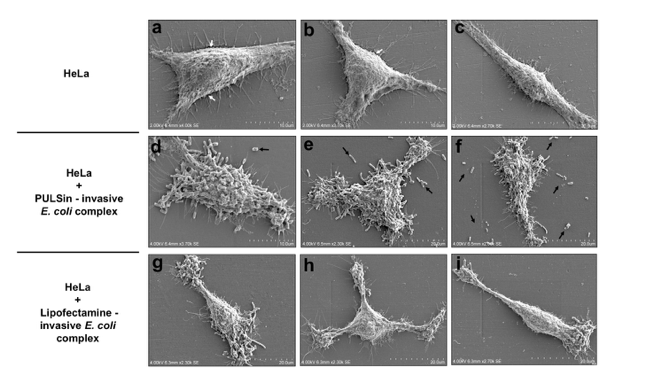
Figure 2: SEM of HeLa with attached E. coli DH10B fixed on plastic coverslips and dehydrated using EtOH gradients without the use of osmium tetroxide or critical point/freeze drying. (abc) HeLa cells allowed to attach overnight, (def) HeLa cells treated with 1 mg PULSin – invasive E. coli DH10B complex at an MOI of 1000 showing diffused adherence patterns and (ghi) HeLa cells treated with 1 mg lipofectamine – invasive E. coli DH10B at an MOI of 1000 showing polarized adherence. Republished with permission from Wiley from: https://doi.org/10.1111/j.1365-2818.1976.tb01075.x
This work was conducted by A. Osahor, K. Deekonda, and K. Narayanan from Monash University Malaysia, C. W. Lee from the University of Malaya, E. U. Sim from Universiti Malaysia Sarawak, and A. Radu from the Icahn School of Medicine at Mount Sinai-New York. This work was funded by a MOSTI eScience grant 02-02-10- SF0252 from the Ministry of Science, Technology and Innovation, Malaysia to K.N.








