
Since the time hydrogen peroxide has been introduced in the markets, there has been a steady increase in the things this wonder chemical can do. From being used as a mild disinfectant, antibiotic, hair bleaching agent, instant fix for tooth whitening, adding the desired highlights to your hair, to being used in cleaning tiles, toilet bowls, rugs and carpets, and even drying acne.
There seems to be nothing that hydrogen peroxide (H2O2, simple peroxide i.e. a compound having two oxygen atoms attached by a covalent single bond H-O-O-H) can’t do to ease your day-to-day life. But there are some contamination issues in the squeaky-clean laboratory setting that has even scientists scratching their heads to find answers for.
One of such pestering issues is the contamination of nucleases which ruin the quality of the nucleic acids (DNA and RNA) isolated from various precious samples. Nucleases especially the ones that can degrade RNA (RNAse A) is so sturdy that one can’t get rid of them even by boiling. This kind of contamination not only compromises the quality of the genetic material isolated but also prevents the scientists from using this isolated material for subsequent high throughput experiments. Researchers use stringent precautions to maintain sterile conditions when they isolate RNA, nevertheless it is not uncommon to find samples degraded.
In a recent report published in a leading scientific journal, scientists used hydrogen peroxide to successfully defeat the menace of nucleases that might still be active in the agarose or running buffer which they use to qualitatively analyses the quality of the genetic material. Several methods are already established which can achieve the desired results. But, they either use noxious chemicals like methylmercury hydroxide, formaldehyde, formamide (potent carcinogens which not only expose the lab workers to dangerous fumes but their discarding procedures are also pretty lengthy) and urea for rather complicated instrumentation.
This recent research has simplified it by adding the humble hydrogen peroxide. According to the scientists, 1ml of commercial hydrogen peroxide (30%w/v conc.) is added to the 100 ml mixture of agarose and buffer and allowed to sit for about 10 minutes. Subsequently, the entire mixture is boiled to melt the agar and casted in the electrophoresis cassettes, and it efficiently rids of all the nucleases. This method is not only quick, it is environment-friendly too. Boiling of the hydrogen peroxide mixture basically breaks the remaining H2O2 molecules into water and oxygen, hence being safe to the researchers and easy to discard.
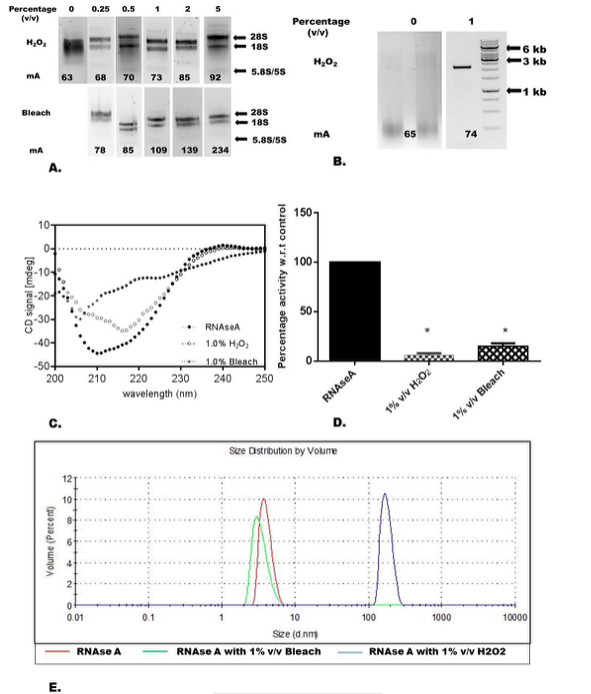
Image republished with permission from Elsevier
The paper further goes on to explain the mechanism by which hydrogen peroxide achieves this amazing feat. Hydrogen peroxide unleashes a two-pronged attack on nucleases. Firstly, by altering the secondary structure of the nuclease enzyme (i.e. it affects the way a protein folds hence rendering it ineffective) and secondly by causing the proteins to aggregate together and become too crowded to be functional. Nucleases are proteins and proteins work best when they are structurally in the correct conformation and exist either as a single entity or usually in a dimer or trimer conformation. When either the structural backbone of the protein is altered or when misfolded proteins accumulate or clump together their activity is either completely lost or severy compromised.
Hydrogen peroxide-based effective inactivation of RNAse A (an RNA nuclease) and DNAse (a DNA nuclease) is indeed a novel, safe, eco-friendly method to reduce the cost on laboratory funds, the health of the researchers and the environment.
These findings are described in the article entitled Hydrogen peroxide agarose gels for electrophoretic analysis of RNA, recently published in the journal Analytical Biochemistry. This work was conducted by Renu Pandey and Daman Saluja from the University of Delhi.
Reference:
- Renu Pandey, Daman Saluja. Hydrogen peroxide agarose gels for electrophoretic analysis of RNA, Analytical Biochemistry, Volume 534, 2017, Pages 24-27, ISSN 0003-2697, https://linkinghub.elsevier.com/retrieve/pii/S0003269717302919.








