
Though DNA has traditionally been depicted as a double helix, a new study published in the journal Nature Chemistry may have just discovered a brand new kind of DNA structure, quite unlike the classic double helix figure.
The DNA that was discovered by the team of researchers has been described as a “twisted knot”, a complex structure made out of the same chemical bases as regular DNA but arrayed in a structure that is much more complex and intricate.
The Composition Of The Twisted DNA Structure
DNA is composed of four different chemical bases: adenine, guanine, thymine, and cytosine. These bases link together to create the famous double helix structure. Yet scientists have theorized that there could be other structures created out of the bases. The new study is the first direct evidence that these alternatives structures may exist within human cells, and that they may play a pivotal role in the regulation of our genome.
The bases are arrayed in an “intercalated motif” structure, an i-structure.
“The i-motif is a four-stranded ‘knot’ of DNA,” explained Marcel Dinger, from the Garvan Institute for Medical Research in Australia, one of the lead authors on the study.
Dinger says that in the knot’s structure the C (cytosine) bases are capable of binding to one another, leading to a situation where the DNA seems to branch outward in a structure made purely of C’s. This is much different from the standard double helix, where C bases bind to G bases (guanine).
Finding The Elusive I-Structure
The structure had been discovered by researchers back in the 1990s, yet because of its unique shape and the acidic conditions that were required to synthesize it, it had only ever been produced in a lab. It was thought that the i-structure couldn’t manifest itself naturally within human cells, but more recent research into the structure found that it was capable of manifesting in less acidic environments.
This new research inspired Dinger and colleagues to see if the i-structure DNA could potentially be found in human cells. Dinger worked alongside Daniel Christ, an antibody therapeutics research from the Garvan Institute to develop an antibody capable of finding any i-structures and binding to them. The antibodies were created with a special chemical marker that would make them glow under certain fluorescent lights.
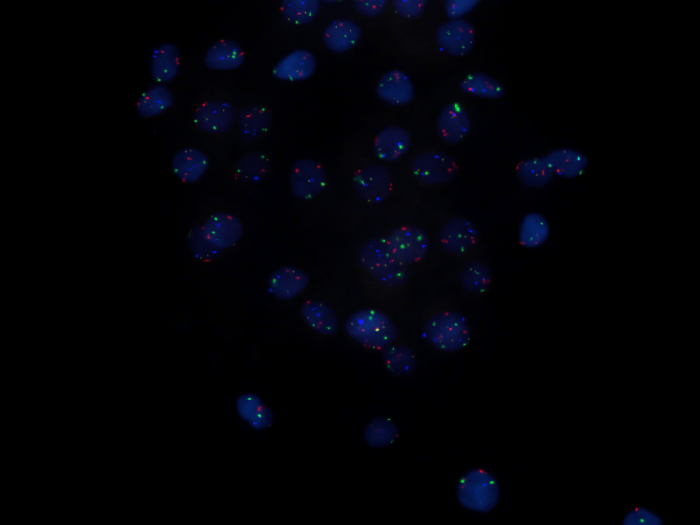
Scientists often stain or tag parts of cells to help them find cerain structures. In this case regions of Urothelial cells were stained by DAPI and tagged with FISH probes. Zeraati’s team also used a chemical marker than would glow under light. Photo: By Bioview – Bioview Duet System, CC BY-SA 3.0, https://commons.wikimedia.org/w/index.php?curid=2513567
By analyzing where the glowing spots were in the DNA sample, they would be able to determine if any i-structure were present and where they were located. Mahdi Zeraati, first author on the study, says that their research suggests the manifestation and disappearance of i-structures is part of a cell’s normal operation.
What excited us most is that we could see the green spots – the i-motifs – appearing and disappearing over time, so we know that they are forming, dissolving and forming again.
Dinger added that it was very likely that our genomes cause all our cells to form i-motifs at some point in their life cycle. Yet these structures disappear very quickly, and the short period of time they are around may explain why they’ve been so hard to detect.
What Function Does The I-Structure Have?
The exact function of the i-motifs is one of the next major mysteries to unravel. The researchers have theorized a possible function for the i-structure; it may help control what genes are expressed.
I-motifs are “dynamic” in nature, meaning that they are capable of unfolding and refolding based upon the pH (acidic) of their environment. The sequences of DNA that create i-motifs aren’t usually found within the genes themselves, rather i-motifs usually occur at “promoter regions”. Promoter regions are sections of the DNA that control which genes are turned on or off, expressed or not expressed. I-motifs were also found within telomeres, a type of genetic marker that influences aging. The research team says that it seems likely that the i-motif assist genes with turning on and off, and they may affect if certain genes are actively read.
Various cellular stressors could be responsible for making the pH inside the cell change, leading to the formation of the i-motif. This change could then alter how a nearby gene is expressed. It could cause the gene to be over-expressed or under-expressed, right now there’s no way of knowing for sure.
Randy Wadkins, a biochemist from the University of Mississippi, was not involved with the study but he warns against being too quick to assume that the i-structure has some great significance. Wadkins says it’s possible that the i-structures may have simply been caught by the antibodies as they were forming, and that they don’t have any real biological significance. In other words, the structure could be an anomaly that forms for a second and then is quickly dissolved, having no real impact on the biology of the cell overall. DNA is, after all, a “conformationally flexible molecule”, and it may take on all kinds of weird shapes that don’t serve biological functions.
Other Alternative DNA Structures?
Indeed, the i-structure isn’t the only alternative DNA structure that might exist in human cells. Scientists have also succeeded in synthesizing a variety of different DNA structures such as A-DNA, B-DNA, Z-DNA, G quadruplex, and triplex DNA. Most of these alternative DNA structures have only been found in laboratory conditions, though the G quadruplex structure was in human cells somewhat recently.
It’s unknown how many of the other DNA structures might appear in the human cell, but now that scientists have found instances of the i-structure in human cells, research into the other structures seems likely. For now, the research team is set to continue investigating the i-structure in attempts to discern what, if any, function it has. The research team is preparing for a “new push to understand what this new DNA shape is really for” and to understand if it will have an impact on health and disease research.









