
Bacteria are known for their ability to rapidly evolve and respond to new threats, which is why drug-resistant bacteria have become such a threat to public health. However, recent research into bacterial evolution mechanisms has revealed one of the mechanisms responsible for their adaptability.
A study recently published in the journal Nature Microbiology reports that bacteria have the ability to harpoon sections of DNA in the environment around them, incorporating them into their body in order to speed up evolution.
Confirming A Long-Held Theory
Researchers were studying Vibrio cholerae bacteria, which is the pathogen responsible for the disease called cholera. The researchers managed to catch on camera a long tendril shooting out from one of the bacterium, spearing a piece of DNA and reeling it back to its body where it absorbs it. The appendage is known as a pili, and the DNA that the bacteria collected with it is incorporated into its own DNA with a process known as horizontal gene transfer. Horizontal gene transfer is distinct from vertical gene transfer, which is the passing of genes down from parent to offspring. It has been hypothesized for decades that bacteria may use a pilus to facilitate gene transfer, but this is the first time that scientists have ever witnessed the event occurring.
Biologist Ankur Dalia, from Indiana University Bloomington, explains that the technique of horizontal gene transfer is an extremely important mechanism to study because it’s likely how antibiotic resistance can move between bacterial species. The structures involved with this process are incredibly small, the width of a pilus is 10,000 times smaller than the width of a human hair, which is why this is the first time the process was ever documented. The verification of this theory is important, as it may drive efforts to better understand the process, which increases the chance that scientists could discover a way to thwart this process and fight antibiotic-resistant bacteria.
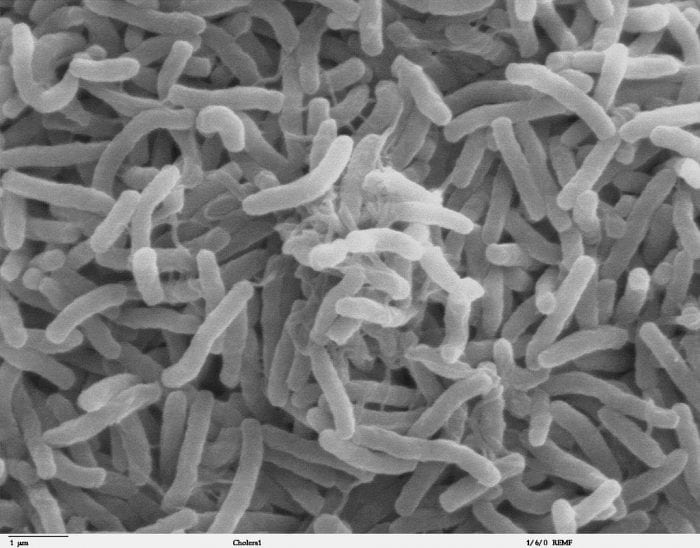
Photo: Copyrighted free use, https://commons.wikimedia.org/w/index.php?curid=197609
Antibiotic resistance has quickly become one of the primary health concerns of the 21st century. The World Health Organization estimates that every year approximately 1 million people are affected by some form of antibiotic-resistant bacteria. This includes around half a million people affected with antibiotic-resistant tuberculosis and another half million people infected with other resistant diseases. The Center for Disease Control reports that approximately 23,000 deaths have occurred within the United States due to bacteria resistant to antibiotics.
The technique that allowed the researchers to document the phenomena is a new system of staining. The staining/painting method allowed researchers to ink both the DNA and the pili with a dye that fluoresces under the right conditions. Then when they stuck the dyed samples under the microscope, they could, for the first time ever, see the pili-gene transfer process occur.
Spearing DNA
Close observation of the bacteria in the pili involved documented how the V. cholera bacteria were able to latch onto chunks of DNA with the tip of the pili then subsequently hold and pull the DNA through a tiny pore in the surface of their bodies. They seem to use some sort of ratcheting mechanism to accomplish this. The pili of V. Cholerae are capable of extending about as far as the length of the bacteria itself. The bacteria can create between 1 to 2 of them every minute, which they retract and extend with a series of proteins that are created and then quickly disassembled.
The pili used by bacteria are capable of piercing the wall of the cell and connecting to chunks of DNA, which can then be drawn out of the cell and back to the bacteria with incredibly precise movements. The amount of precision required for bacteria to collect DNA is incredible, according to biologist Courtney Ellison who describes the process like “threading a needle”.
The plasma membrane of a bacterium is made out of a phospholipid bilayer. These are two layers of lipid molecules arrayed in opposite directions. There is a phosphorus head and a fatty acid tail. Because of the bilayer’s construction, the contents of the cell are kept inside and separated from the outside environment. The membrane is selectively permeable and large molecules (like DNA) can only move into the cell at special pores, or openings within the bilayer.
According to Ellison the width of the whole found in the outer membrane of the cell is just large enough to pull a DNA helix bent in half out from the cell. If a pilus wasn’t guiding it, it’s unlikely the DNA could be brought into the cell. Explains Ellison:
“The size of the hole in the outer membrane is almost the exact width of a DNA helix bent in half, which is likely what is coming across. If there weren’t a pilus to guide it, the chance the DNA would hit the pore at just the right angle to pass into the cell is basically zero.”
This isn’t the only method that bacteria use to gain antibiotic resistance, as resistance to antibiotics can be transferred between generations of bacteria in a few different ways. Horizontal gene transfer can also be accomplished via a few different mechanisms. However, it is still an incredibly important method to understand. In this case, the absorption of DNA present in the bacteria’s immediate environment is referred to as transformation. Transformation is used by the bacteria to collect DNA from other dead bacteria.
Future Implications
When a bacteria dies its membrane splits apart and the DNA within it spills into the environment. When this happens, other bacteria nearby can then fish for this DNA and incorporated within themselves. If the bacteria that died had a resistance to a certain antibiotic, it’s DNA could be collected by other bacteria to grant themselves that resistance and pass that resistance on to their offspring. Because of this process, antibiotic resistance can quickly be distributed out to an entire population of bacteria.
Researchers are evidently interested in discerning the mechanisms that bacteria use to evolve resistances to the antibiotics, in the hopes that they can create ways to circumvent these resistances. Now that researchers know bacteria can indeed harpoon pieces of DNA, they want to find out how the bacteria’s pili can anchor to DNA at the correct place. As mentioned, precision is necessary for this operation to succeed. The proteins that bacteria use for this process appear to have interactions with DNA that haven’t been witnessed before.
The research team hopes that the method of applying fluorescent dye they use to observe the pili fishing for DNA can help them determine the other functions of the pili.
“These are really versatile appendages,” explained Dalia. “This method invented at IU is really opening up our basic understanding about a whole range of bacterial functions.”









