
Human life has always depended on natural resources. Nature provides our housing, food, fragrances, and oils, and is our pharmacy. It remains the basis of health care for 80% of the world’s population who rely on traditional medicines to treat disease or promote health. The pharmaceutical industry continues to find in natural products an unrivaled source of chemical and biological inspiration.
Pharmacognosy is defined as the study of biologically active natural products and their uses by humans for healing purposes. Different societies have access to different methods to gain insight into the effects of natural products (see Fig. 1). These methods can range from consuming psychotropic “teacher plants” to the query of in silico fragmented databases established through machine learning. Despite their differences, both approaches, and many others are valid and often essential to the overall health of the society of which they are a part.
This presentation pleads for a global vision of this discipline. We believe that combining the different approaches for integrated knowledge acquisition and assessment can only enhance human global health in a sustainable manner. With this commitment, we present a series of challenges, the major one consisting in developing an efficient system for knowledge acquisition and sharing across time, geography and interested parties. (1)
Pharmacognosy is an intensely data-rich natural science founded in taxonomy, chemistry, and biology. Despite the ever-increasing volume of data being acquired, most are not widely accessible and even less properly curated, annotated, or qualified. As a result, the utility of this vast store of anecdotal and experimental information has not been effectively assessed and potentiated. Collections of large datasets are frequently found in a single location (a series of facial images or photos, social networking sites, or marketing information, etc.), and are readily subjected to consolidated analytical processing by the company that acquired them or by third-parties. For pharmacognosy, these data sites are dissipated and dispersed as numerous silos worldwide, with essentially no communication among them. Thus, although acutely data-rich, pharmacognosy is information poor. To enhance the role of natural products in a burgeoning society, which has an underpinning of sustainability, pharmacognosy must organize and make its data accessible and globally understandable. According to the Convention on Biological Diversity, this will require full consideration of the issues of intellectual property as well.
For this to be achieved, an efficient database ecosystem should be designed. Natural Products Alert (NAPRALERT) at the University of Illinois at Chicago was the first and is the only, database accumulating, in great detail, ethnomedical, taxonomic, chemical, and biological (in vitro, in vivo, and clinical) information gleaned from the past and contemporary natural product literature. In a digitally-enhanced format, where global knowledge can be acquired from a hand-held device, NAPRALERT should stand as one of the corners in a comprehensive system of natural product databases. For the evolution of an impactful future, the natural product scientist needs a portal, a Global Archive of Natural Products (4), through which searches for many diverse aspects (taxonomy, spectra, network pharmacology, metabolomic analyses, genomics data, etc.) of curated natural product information can be linked to and from other resources.
The ecopharmacognosy concept brings focus to a sustainability philosophy and its associated practices in the development and use of biologically active natural products (2,3). Ecopharmacognosy extends the “green” chemistry concepts to the sustainable provision of natural resources and the knowledge produced from them. The Anthropocene era of massive human development has resulted in incredible benefits for civilization and enormous devastation for the planet. (5) It is now timely to employ high-tech, integrated, information-based solutions in the interest of all life on Earth.
To reduce the use of non-sustainable resources in natural product research, the quality and use of in silico datasets should be improved. Indeed the interrogation of efficiently curated datasets can avoid the tedious and costly reisolation of known natural products. It is estimated that each dereplicated (identified prior to its physical isolation) natural product saves ca. $50,000 (in addition to the saved working time, solvents and biological raw material needed for the study). (6) Other applications include: i) accumulation of genetic data on organisms, and its use for authentication, establishment of taxonomic relationships or gene clusters mining for the obtention of targeted metabolites; ii) metabolomics to assess biosynthetic pathways, and establish acceptable ranges of similarity for chemical verification, particularly with respect to environmental or quality control processing factors; iii) in-field detection of dominant metabolites using non-invasive Raman or infrared spectroscopy techniques; and vi) active site modeling techniques for optimizing potential binding parameters without isolating the natural products and prior to targeted synthesis. It is these integrated datasets which will enhance the efficiency and effectiveness of natural product research, and foster the prioritization of that research for societal benefit.
This productive discipline will progress by mining, enhancing and sharing legacy and future published research and knowledge. It will require a serious and respectful integration of the methods of knowledge acquisition of natural products, which are, for now, external to the classical discipline. This global translational approach should greatly benefit human health.
Republished with permission from Elsevier from https://doi.org/10.1016/j.copbio.2018.02.010
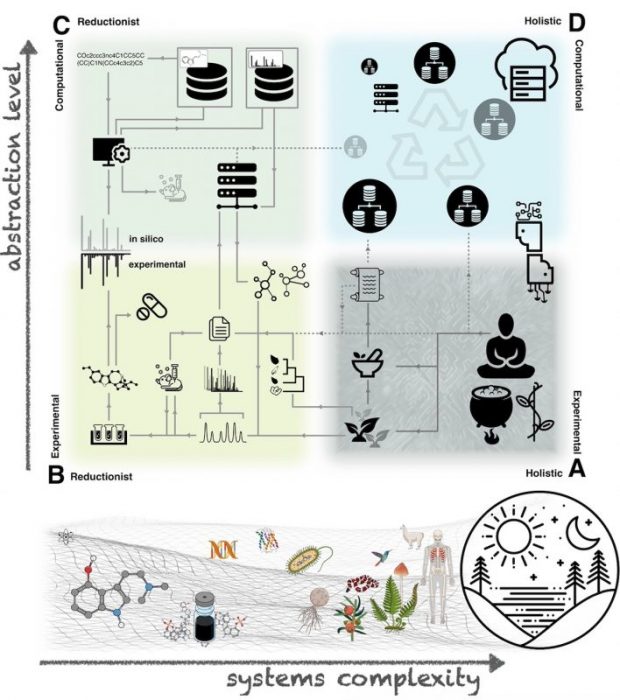
Fig.1 Proposed epistemological framework of knowledge acquisition methods in pharmacognosy. At the bottom are depicted the main subjects of study of the discipline in order of increasing systems complexity: single chemical entities, natural product extracts, genetic material, proteins, microorganisms, plants, animals (including humans), the whole planet ecosystem and their possible interactions. Above are schematized knowledge acquisition methods in the discipline. They were divided four ways according to their tendency to rely on: holistic and experimental approaches (Corner A, e.g. traditional medicines), reductionist and experimental approaches (Corner B, e.g. bio-guided fractionation), reductionist and computational approaches (Corner C, e.g. in silico fragmentation of a single chemical entity) and holistic and computational approaches (Corner D, a hypothetical ecosystem of open databases aggregating pharmacognostic knowledge). Grey links represent examples of information fluxes (solid lines as existing fluxes and dashed lines as potential ones). These knowledge acquisition methods are not mutually exclusive. For example: the use of an in silico annotated dataset about ethnopharmacologically surveyed plants can highlight metabolites that could support their reported traditional use. Such an approach would feed on the three knowledge acquisition methods (Corners A, B, and C). (1)
These findings are described in the article entitled Pharmacognosy in the digital era: shifting to contextualized metabolomics, recently published in the journal Current Opinion in Biotechnology. This work was conducted by Pierre-Marie Allard, Antonio Azzollini, and Jean-Luc Wolfender from the School of Pharmaceutical Sciences, University of Geneva, University of Lausanne, Jonathan Bisson and Guido F Pauli from the University of Illinois at Chicago, and Geoffrey A Cordell from the University of Florida and Natural Products, Inc.
References:
- Allard, P.; Bisson, J.; Azzollini, A.; Pauli, G. F.; Cordell, G. A.; Wolfender, J. Curr. Opin. Biotechnol. 2018, 54, 57–64.
- G.A. Cordell. Ecopharmacognosy – the Responsibilities of Natural Product Research to Sustainability. Phytochem. Lett., 11, 332-346 (2015).
- G.A. Cordell. Cognate and Cognitive Ecopharmacognosy – in an Anthropogenic Era. Phytochem. Lett., 20, 540-549 (2017).
- G.A. Cordell. Sixty Challenges – A 2030 Perspective on Natural Products and Medicines Security. Nat. Prod. Commun.,12, 1371-1379 (2017).
- Anonymous, 2012. State of the Planet Conference. http://stateoftheplanet.org/(accessed 04.07.16[GC3] .).
- Corley, D.G. and Durley, R.C. (1994) Strategies for database dereplication of natural products. J. Nat. Prod. 57, 1484–1490









