Influenza A viruses (IAVs) are well known for annual, seasonal disease outbreaks resulting in mild to severe consequences from economic losses to hospitalization and death. These viruses are often thought of as human viruses, but in fact, an extensive reservoir of influenza A viruses exist in animal populations, such as wild and domesticated birds, pigs, horses, and seals. Only 3 virus subtypes — H1N1, H2N2, and H3N2 — have circulated extensively in humans over the past century to cause seasonal epidemics. Each virus entered the population via a pandemic event that was preceded by adaptation of an animal strain to a naïve human host.
The risk for additional viruses adapting and exhibiting cross-species transmission is of concern, as highly pathogenic strains of H5 and H7 subtypes arise and circulate in avian species at unpredictable intervals. A number of species barriers exist that slow down this process of cross-species adaptation, including structural nuances of the host cell receptor.
Influenza A virus is believed to bind to sialylated glycans (complex carbohydrates) on cell surfaces, which leads to the entry of the virus into cells presumably by the process of endocytosis. Within the endosome at low pH, the viral glycoprotein hemagglutinin (HA) facilitates the fusion of the viral and endosomal membranes, allowing for the release and, ultimately, replication of the IAV genome within the cytoplasm of the infected cell. Sialic acids in mammalian glycans are usually either alpha 2,3 or 2,6 linked to a penultimate galactose, and avian viruses tend to recognize sialic acid in the alpha 2,3 linkage, whereas human and swine viruses prefer an alpha 2,6 linkage.
The existing theory is that this linkage preference corresponds to the presence of each particular type of glycan in the tissue at the site of infection in the host. The receptor binding characteristics of various HAs and whole viruses have been examined in detail; however, each method used (specifically sialylated red blood cells, synthetic glycan microarrays, and biolayer interferometry with short 2,3 or 2,6 sialylated glycans) is limited by the biological significance of the glycans presented. In short, we are able to learn what structural determinants HA can bind but are unable to say if such determinants are present in the tissue at the site of infection and how important such determinants are to the infection of cells.
One reason for this uncertainty is that the glycome of an entire human organ has not been examined in great detail. In our studies, we sought to isolate and separate the glycans from human lung and generate a glycan microarray to probe IAV interactions. We were able to release the glycans from human lung tissue from an organ donation program when a suitable match for transplant was not found, using the ORNG method (Oxidative Release of Natural Glycans). The released glycans were tagged with a bi-functional linker, separated into fractions, and printed covalently on glass slides to prepare the first human lung shotgun glycan microarray (HL-SGM). As we are able to fluorescently label IAV particles, we incubated these viruses on the HL-SGM and then quantified the fluorescence of the spots where the viruses bound to printed glycans. At this point, we could return to the fraction that each printed sample represents and structurally characterize the glycans within.
We discovered that while a variety of viruses (different subtypes, avian, swine and human hosts, and different times of isolation) bind to human lung glycans on the array, they bound to different types of glycans and with different patterns. Some of these human lung glycans were sialylated and resembled those that have been reported in other natural tissues, such as pig lungs, for being important for influenza A virus binding. These included bi-antennary di-sialylated N-glycans. We found that both alpha 2,3 and 2,6-linked sialic acid are present in human lung glycans, especially on the common bi-antennary N-glycan structures, though, due to the nature of the organ processing, we do not know yet whether these linkage types are localized in a distinct part of the lung or colocalized. Most importantly, we found that each virus also exhibits robust binding to a set of unexpected glycans that lack sialic acid — those containing phosphorylated high mannose moieties. Binding competition assays with free sialic acid and sialylactose suggest that the binding to phosphorylated glycans is not occurring at the canonical receptor binding site on the HA and may involve a unique site of interaction.
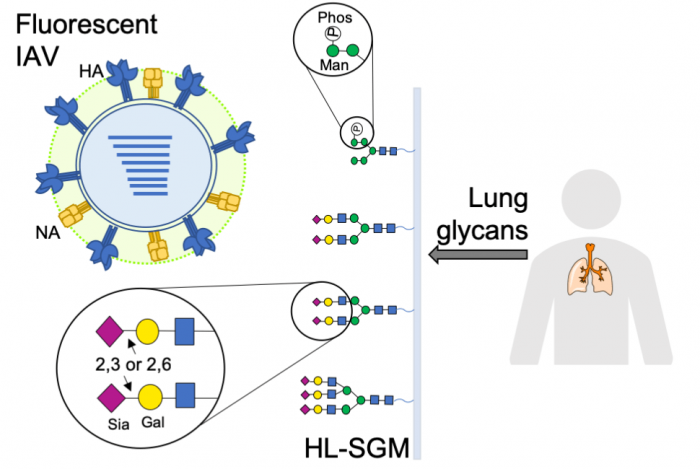
N-glycans are isolated from human lung tissue and printed to generate the human lung glycan microarray (HL-SGM). Fluorescently labeled influenza A viruses (IAV) bind to certain glycans on the array but not others, revealing those that may function as host cell receptors for infection. Important structural determinants include sialylated, branched glycans and unexpectedly, phosphorylated high mannose glycans. Figure courtesy L. Byrd-Leotis
Thus, the generation of the HL-SGM has enabled us to describe a previously unknown binding interaction of IAV with glycans from the human lung, in addition to providing a comprehensive description of the human lung glycome. Though much is still to be learned about the binding of phosphorylated glycans, including the mechanism and function, such a discovery is a paramount first step to considering alternative receptor interactions that could be important in binding, fusion, or the viral assembly process.
This work is the first to utilize the glycome of an entire human organ to better reflect the biological context of virus-host interaction and, along with future comparative glycomics studies, will pave the way for understanding the human glycome. The HL-SGM resource itself will be useful as a tool for pathogen studies beyond influenza A virus interaction. Additionally, as more IAV isolates are obtained that have recently adapted or exhibit so-called pandemic potential, we will be able to provide a snapshot of the potential interactions with lung tissue for infection, shedding light on the process of cross-species adaptation.
These findings are described in the article entitled Influenza binds phosphorylated glycans from human lung, recently published in the journal Science Advances.









