
Although there is still no cure for HIV, HIV-positive patients can live a near normal life expectancy with modern therapeutic regimens. Thanks to affordable diagnostic tests and highly effective antiretroviral therapy, the annual number of deaths due to HIV has been decreasing since 2006.
However, as people live longer with HIV, another problem has gained attention: drug resistance. HIV drugs work by suppressing the amount of virus in a patient’s blood. As HIV replicates, it is prone to accumulating mutations. If these mutations are located in regions acted upon by antiretroviral drugs, therapeutic efficacy can be compromised. Without modification to appropriate second-line treatment, HIV will continue to replicate, and drug-resistant strains can begin to dominate as well as be transferred to others. Unfortunately, rates of resistance are on the rise and are above 10% in some areas. For these reasons, it’s important to catch resistance as soon as possible.
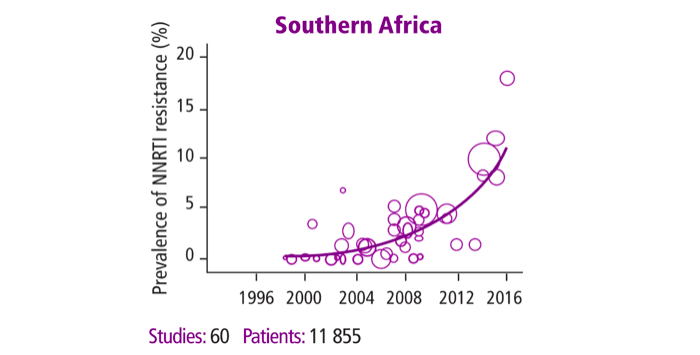
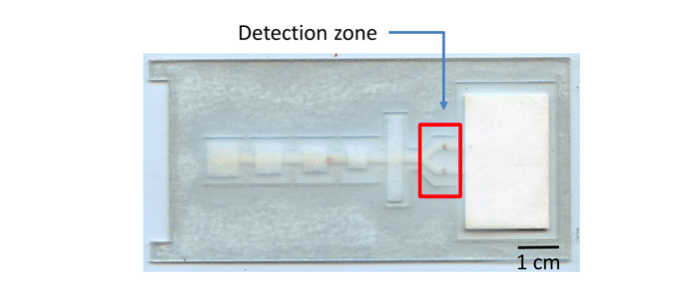
Figure 1: Prevalence of pre-treatment resistance to first-line antiretroviral therapy. HIV Drug Resistance Report, 2017, World Health Organization.
Advanced healthcare systems, such as those in the US, identify drug resistance by sequencing the viral genome and identifying all mutations present. But sequencing requires high-tech laboratory equipment and several weeks, and costs between $200-700. Two-thirds of the 36.7 million people living with HIV/AIDS worldwide reside in sub-Saharan Africa, where access to resistance testing is limited. Lack of infrastructure in these areas has resulted in a reliance on clinical monitoring only or clinical monitoring in combination with immune system monitoring.
Where these luxuries aren’t available, clinicians often either prescribe second-line drugs or keep the patient on ineffective first-line drugs because they suspect nonadherence. Without knowing if the virus has actually mutated, we are wasting money and valuable resources, and allowing HIV drug resistance to spread. Near-point-of-care assays, or tests that can be used in health posts without many resources, are likely to be the most useful option to detect key drug resistance mutations in the most remote settings.
To address this issue, a group based at Rice University is working to increase access to resistance testing. A recent paper published in Analytical Biochemistry describes the development of a cheap and disposable device that will discriminate between mutant and naturally occurring HIV. The project aims to amplify DNA to detectable levels and visually display results on a strip similar to a pregnancy test – all for under $13. Clinicians could then make an immediate decision on patient treatment.
Instead of sequencing the viral genome, the Rice group chose to focus on a section of the HIV genome that contains several known high-profile mutations. By amplifying this section of the genome, 90% of patients with resistance would be identified. The first step of the assay consists of nucleic acid amplification. Instead of using polymerase chain reaction, which requires thermal cycling between high and low temperatures to denature and anneal nucleic acids, the paper uses recombinase polymerase amplification, a method that uses a combination of enzymes and proteins to perform amplification isothermally at 37C. Next, the authors tag wild-type and mutant HIV DNA with different moieties for detection using a thermal cycler, while future work will focus on performing this step isothermally.
To detect the products, the group built on previous work from their lab and others to develop a paper and plastic enzyme-linked immunosorbent assay (ELISA). Traditional ELISA reagents are added to glass fiber pads, along with the sample and buffers to promote fluid flow. The device is constructed on a backing of adhesive clear acetate. The test is folded across the red vertical line and sealed shut, and a cellulose wicking pad drives fluid flow through a nitrocellulose membrane through capillary action.
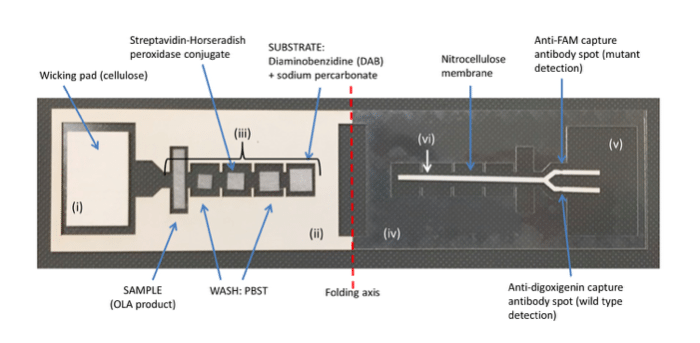
Figure 2: Image of the lateral flow ELISA to discriminate HIV-1 mutant and wild-type OLA products. After reagents are added, the adhesive backing on the left is removed and the device is folded and sealed along the folding axis to initiate flow and sequential delivery of reagents. The device parts indicated are: (i) cellulose wicking pad; (ii) adhesive acetate (paper side up); (iii) glass fiber pads; (iv) acetate; (v) adhesive acetate (paper backing removed and adhesive side up); (vi) nitrocellulose.
After 45 minutes, the results of the test are displayed in a visual format, with one dark brown spot appearing to indicate wild-type DNA, and another appearing to indicate the presence of mutant DNA. Although the test can easily be read by eye, the authors hope to incorporate a handheld reader in the future. The authors developed this test for the most common HIV drug resistance mutation, M184V, but the assay could be easily adapted for other known mutations.
These findings are described in the article entitled Paper-based detection of HIV-1 drug resistance using isothermal amplification and an oligonucleotide ligation assay, recently published in the journal Analytical Biochemistry. This work was conducted by Mary E. Natoli, Carolina De Santiago, and Rebecca R. Richards-Kortum from Rice University, Brittany A. Rohrman from Omniome, and Gert U. van Zyl from Stellenbosch University.









