
Published by Anu Raghunathan
Chemical Engineering Division, CSIR-National Chemical Laboratory, Pune, Maharashtra, India
These findings are described in the article entitled Constraints-based analysis identifies NAD+ recycling through metabolic reprogramming in antibiotic resistant Chromobacterium violaceum, recently published in the journal PLoS One (PLoS ONE 14(1): e0210008). This work was conducted by Deepanwita Banerjee and Anu Raghunathan from CSIR-National Chemical Laboratory.
The leading edge of the global problem of antibiotic resistance necessitates novel therapeutic strategies (BMC Syst Biol. 2017; 11: 51). Infections, in the 20th century, have been considered to no longer be life-threatening since antibiotics were discovered. However, antimicrobial resistant pathogens are projected to cause death in more people than cancer by 2050. While there are disputes about the projected death rate and mechanisms of emergence of antibiotic resistance, the fact is that bacteria are evolving resistance to antibiotics faster than the drug pipeline is being replenished. In this war against antimicrobial resistance, researchers are trying desperately to develop new drugs or slow down emerging resistance. One recent approach has been to extend the shelf life of antibiotics as it is inevitable that new drugs will eventually be rendered ineffective on these superbugs.
So what is antibiotic resistance? There are many definitions and in the simplest terms, it is the uncontrolled growth of pathogens even in the presence of drugs, at concentrations that are generally therapeutic. So as a corollary, antibiotic resistance generally occurs when the drug has lost its ability to kill or retard pathogen growth. Our laboratory, the Metabolic Inquiry and Cell Engineering lab at CSIR-National Chemical Laboratory, Pune, India tackles this problem by re-sensitizing antibiotic-resistant pathogens (Figure 1) using a model-driven/model integrated systems approach and identifying targetable metabolic reactions and pathways that can kill these superbugs.
The two principal dimensions of life or cell function are energy generation and growth. Our publication explores the duality of cell function, primarily orchestrated by metabolic networks, critical for survival and hence governing resistance. The most exciting aspect of this study is the computational ability to analyze the differences in the fundamental metabolism of antibiotic-resistant pathogens and predict the rewiring of metabolic networks that need to be reversed to kill resistant superbugs.
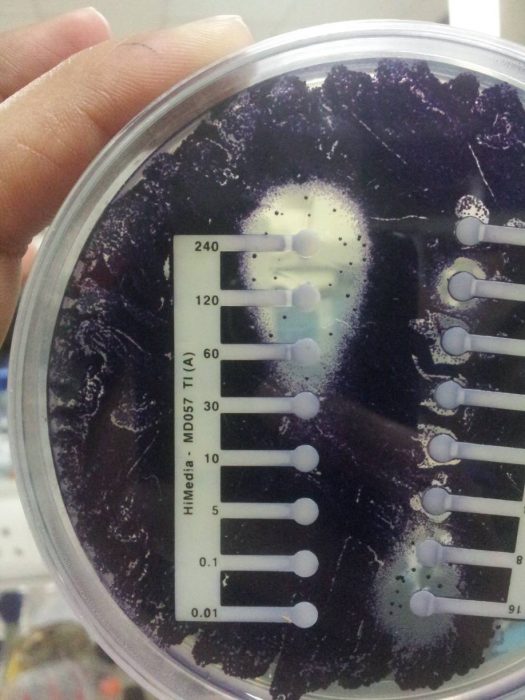
Figure 1: The evolved antibiotic resistant Chromobacterium violaceum.
Photograph Credit: Dr. Deepanwita Banerjee
In our recent Publication in PLOS One, we have developed a genome-scale metabolic network reconstruction of a zoonotic opportunistic pathogen, Chromobacterium violaceum and translated it into a mathematical model. A technique called constraints-based flux balance modeling is then used to compute cell function/phenotype. These methods are scalable and can be extended to any infection causing pathogen.
C violaceumis a gram-negative bacterium, purple in color with a metallic sheen, due to a pigment it produces called violacein. We have shown in a previous publication (BMC Syst Biol. 2017; 11: 51) that the modulation of the secretion of this pigment can be a potential biomarker for resistance. We use the secretion profiles of this biomarker to define constraints that are used to define antibiotic resistance in our in silicopathogen in this publication. The in silicoorganism (i.e., the genome-scale metabolic model) predicts the impact of Streptomycin and Chloramphenicol, two antibiotics that target protein synthesis, on nutrient utilization, central metabolism, respiration and growth (Figure 2). We identify disruption of central metabolism and rewiring of several reactions using the electron carrier NADH. Disruption of redox homeostasis through NADH/NAD ratios was identified as central to antibiotic action. The electrons from NADH flow through the electron transport chain and cause a build-up of protons across the membrane. This proton gradient or Proton Motive Force (PMF) causes a protein ATP synthase to make ATP. Adenosine Triphosphate (ATP) is the energy currency or store of the cell. Modulated ETC pathway fluxes eventually create differential PMF/ electrochemical membrane gradient for ATP synthesis were identified in the presence of the antibiotics.
The resistant superbug models predicted metabolic reprogramming to compensate for the redox imbalance and differential PMF triggered by the antibiotics. Calculating operational metabolic flux distributions identified reactions that need to carry flux forced in any particular direction for optimal growth to be redirected during suboptimal growth on malate, succinate and pyruvate. This results in failure to reconcile cofactor balance and maintain a rigid core of flux distribution. This would eventually prevent minimal cellular function of the metabolic network for energy and biomass formation.
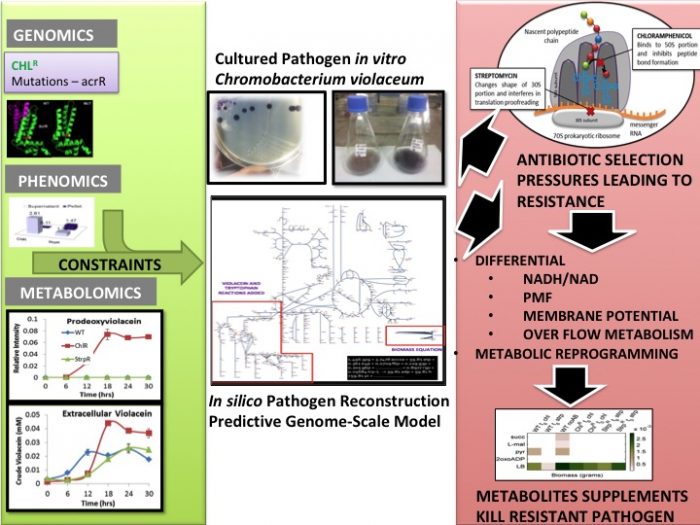
Figure 2: Scalable Methodology to predict metabolic vulnerability: The left-hand panel in green shows experimental data that define constraints of the pathogen in vitro and in silico used in the Genome-Scale Metabolic Model (Middle panel). The right-hand panel in red shows the outcome of antibiotic selection pressures and how they impact different physiological factors that can be integrated into the model to predict the subset of metabolites that would kill the resistant pathogens. Credit: Credit: Dr. Deepanwita Banerjee
Taken together, our data unraveled that disruption of redox homeostasis by certain benign metabolites (Figure 2) as key to killing antibiotic-resistant pathogens. Reversing the phenotype by perturbing metabolism through modulating micro-environments could be one way of subverting the onset of the post-antibiotic era. This could lead to scalable pipelines integrating growth/metabolite/MIC profiling and constraints-based flux balance models for clinical isolates, ultimately leading to personalized treatment and individualized therapy.









